Color residues jmol3/2/2023  
In Jmol, temperature does the same as relativeTemperature. RasMol/Chime note: In RasMol and Chime, temperature (the only option available) uses also a relative scale, but with a reverse rainbow gradient (bgyor). Note: The temperature factor fields in a PDB file are sometimes used by the file creator to indicate any other characteristic of the atoms in the model. RelativeTemperature after set rangeSelected on: the color is relative to the lowest and highest B factor values within the currently selected part of the model. RelativeTemperature: the color is relative to the lowest and highest B factor values within the file (not within the selection). (red = hot = high temperature = variable/mobile)įixedTemperature: the factor is made relative to an absolute scale of 0 to 100. (blue = cold = low temperature = static/immobile) A high crystallographic B factor may indicate an incorrect structure. Typically this gives a measure of the mobility or uncertainty of a given atom's position. Related commands: color object relativeTemperature, color object fixedTemperatureĬolors each atom, using a bwr gradient, according to the "temperature factor" (B factor, or Debye-Waller factor) value stored in the PDB or mmCIF file. If you want to exclude "hetero" groups, then do not select them. In addition, the set hetero command is not implemented in Jmol. RasMol/Chime note: In RasMol/Chime, group index is relative to the full chain, while in Jmol it is relative to the currently selected groups of the chain.  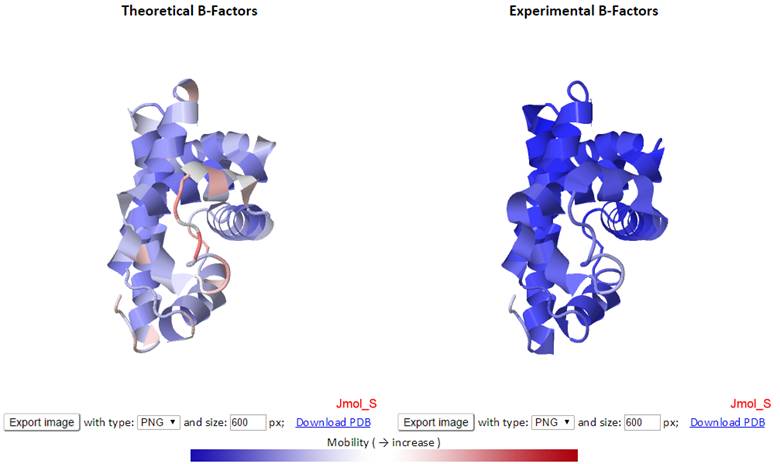
HETERO groups may participate in a polymer, depending upon how the PDB/mmCIF file is marked up, and as long as they have a backbone pattern that Jmol can recognize as a sequence of individual units.Again, the numbering of groups in the file is irrelevant.If a chain has a break in it, the 2 pieces of the chain are recognized as separate polymers, and hence each one is colored independently with the full color range, despite the fact that both parts are designated with the same chain ID in the coordinate file.Jmol interprets the backbone pattern of consecutive residues ("monomers") to identify continuous "polymers".A "polymer" is a series of connected "monomers" (at least for these purposes). The coloring operates independently for each chain.Ī variation, relating to "monomers" in a "polymer" rather than "groups" in a "chain".The order of groups in the file is used, not the number assigned to them in the file.The full color range is applied across the specified or selected groups in the chain (that is, the scale is relative, not absolute).(red = oxygen = C-terminal = end of chain = 3') (blue = nitrogen = N-terminal = start of chain = 5') Backbone displays (such as ribbons, cartoons, etc.) are rendered in the color of alpha carbons for proteins, phosphorus for nucleic acids.ĭefault element colors, by periodic table: Hover over any element to see more information.Ĭlick on an element to move to the same information in the Atomic Numbers table below. Related commands: color object cpk, set defaultColors Jmol, set defaultColors RasmolĪpplies color to each atom of the object according to element, as shown in the tables below. Atoms ('CPK' colors, default element colors) Many of the following coloring patterns apply only to PDB and mmCIF files for biomacromolecules. Information on RasMol colors is included for those who are adapting RasMol and Chime-based materials for use in Jmol. if the object is omitted, atoms is assumed.object can be atoms, bonds, backbone, cartoon, stars, rockets, ribbon, dots, label, echo, hbonds, ssbonds, axes, boundbox, measure, polyhedra, isosurface, pmesh, unitcell.color scheme is specified using pre-defined terms detailed on this page.(For the default colors and color values used in Jmol, see table below.) It can also be a JavaScript color name, a RasMol color name, or a Jmol color name. color is specified as decimal or hexadecimal triplets.Objects can be custom colored in Jmol using the color (or colour) command: color object color or color scheme 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
